None
Note
This tutorial was generated from an IPython notebook that can be accessed from github.
Create 3D boolean masks
In this tutorial we will show how to create 3D boolean masks for
arbitrary latitude and longitude grids. It uses the same algorithm to
determine if a gridpoint is in a region as for the 2D mask. However, it
returns a xarray.Dataset with shape region x lat x lon,
gridpoints that do not fall in a region are False, the gridpoints
that fall in a region are True.
3D masks are convenient as they can be used to directly calculate weighted regional means (over all regions) using xarray v0.15.1 or later. Further, the mask includes the region names and abbreviations as non-dimension coordinates.
Note
3D boolean masks are more convenient to work with but 2D masks may offer some speed gains when calculating regional means. See #226 how weighted regional averages can be calculated with 2D integer mask.
Import regionmask and check the version:
import regionmask
regionmask.__version__
'0.12.1'
Load xarray and numpy:
import numpy as np
import xarray as xr
# don't expand data
xr.set_options(display_style="text", display_expand_data=False, display_width=60)
<xarray.core.options.set_options at 0x7fa922d69160>
Creating a mask
Define a lon/ lat grid with a 1° grid spacing, where the points define the center of the grid:
lon = np.arange(-179.5, 180)
lat = np.arange(-89.5, 90)
We will create a mask with the SREX regions (Seneviratne et al., 2012).
regionmask.defined_regions.srex
<regionmask.Regions 'SREX'>
Source: Seneviratne et al., 2012 (https://www.ipcc.ch/site/assets/uploads/2...
overlap: False
Regions:
1 ALA Alaska/N.W. Canada
2 CGI Canada/Greenl./Icel.
3 WNA W. North America
4 CNA C. North America
5 ENA E. North America
.. .. ...
22 EAS E. Asia
23 SAS S. Asia
24 SEA S.E. Asia
25 NAU N. Australia
26 SAU S. Australia/New Zealand
[26 regions]
The function mask_3D determines which gripoints lie within the
polygon making up each region:
mask = regionmask.defined_regions.srex.mask_3D(lon, lat)
mask
<xarray.DataArray 'mask' (region: 26, lat: 180, lon: 360)> Size: 2MB
False False False False False ... False False False False
Coordinates:
* lat (lat) float64 1kB -89.5 -88.5 ... 88.5 89.5
* lon (lon) float64 3kB -179.5 -178.5 ... 178.5 179.5
* region (region) int64 208B 1 2 3 4 5 ... 23 24 25 26
abbrevs (region) <U3 312B 'ALA' 'CGI' ... 'NAU' 'SAU'
names (region) <U24 2kB 'Alaska/N.W. Canada' ... '...
Attributes:
standard_name: regionAs mentioned, mask is a boolean xarray.Dataset with shape
region x lat x lon. It contains region (=numbers) as
dimension coordinate as well as abbrevs and names as
non-dimension coordinates (see the xarray docs for the details on the
terminology).
Plotting
Plotting individual layers
The four first layers look as follows:
import cartopy.crs as ccrs
import matplotlib.pyplot as plt
from matplotlib import colors as mplc
cmap1 = mplc.ListedColormap(["none", "#9ecae1"])
fg = mask.isel(region=slice(4)).plot(
subplot_kws=dict(projection=ccrs.PlateCarree()),
col="region",
col_wrap=2,
transform=ccrs.PlateCarree(),
add_colorbar=False,
aspect=1.5,
cmap=cmap1,
)
for ax in fg.axes.flatten():
ax.coastlines()
fg.fig.subplots_adjust(hspace=0, wspace=0.1);
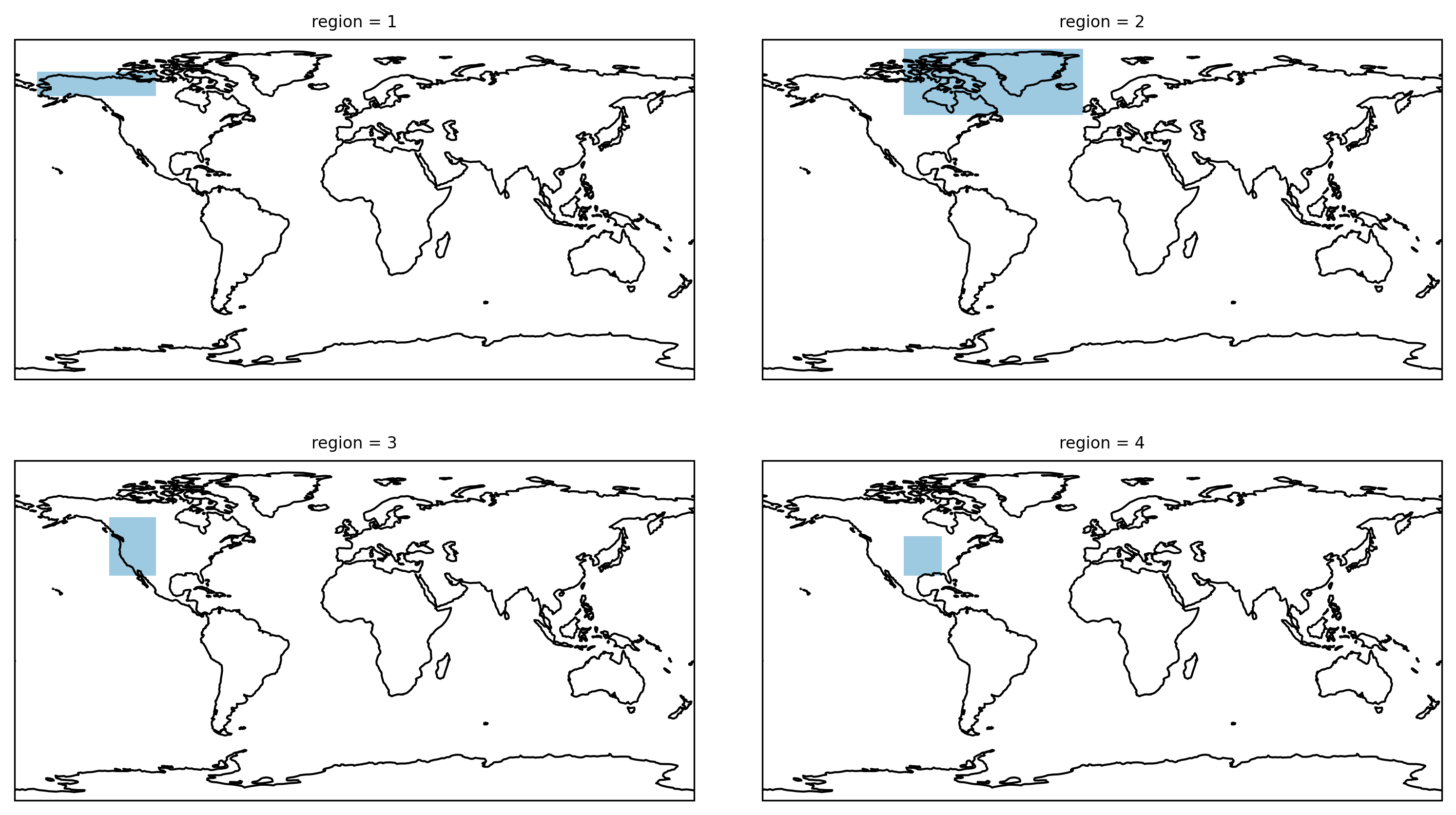
Plotting flattened masks
A 3D mask cannot be directly plotted - it needs to be flattened first.
To do this regionmask offers a convenience function:
regionmask.plot_3D_mask. The function takes a 3D mask as argument,
all other keyword arguments are passed through to
xr.plot.pcolormesh.
regionmask.plot_3D_mask(mask, add_colorbar=False, cmap="plasma");
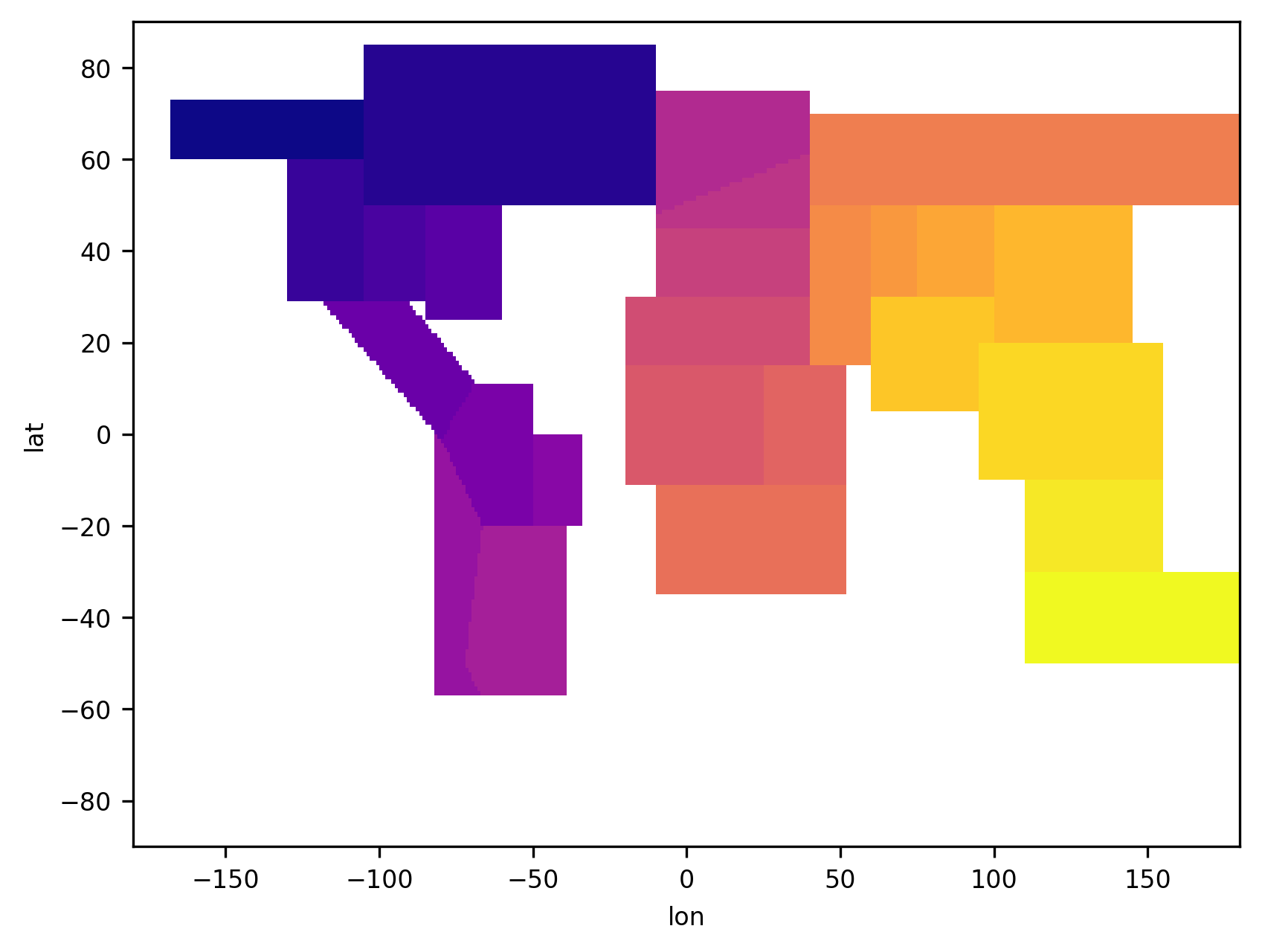
Working with a 3D mask
masks can be used to select data in a certain region and to calculate regional averages - let’s illustrate this with a ‘real’ dataset:
airtemps = xr.tutorial.load_dataset("air_temperature")
The example data is a temperature field over North America. Let’s plot the first time step:
# choose a good projection for regional maps
proj = ccrs.LambertConformal(central_longitude=-100)
ax = plt.subplot(111, projection=proj)
airtemps.isel(time=1).air.plot.pcolormesh(ax=ax, transform=ccrs.PlateCarree())
ax.coastlines();
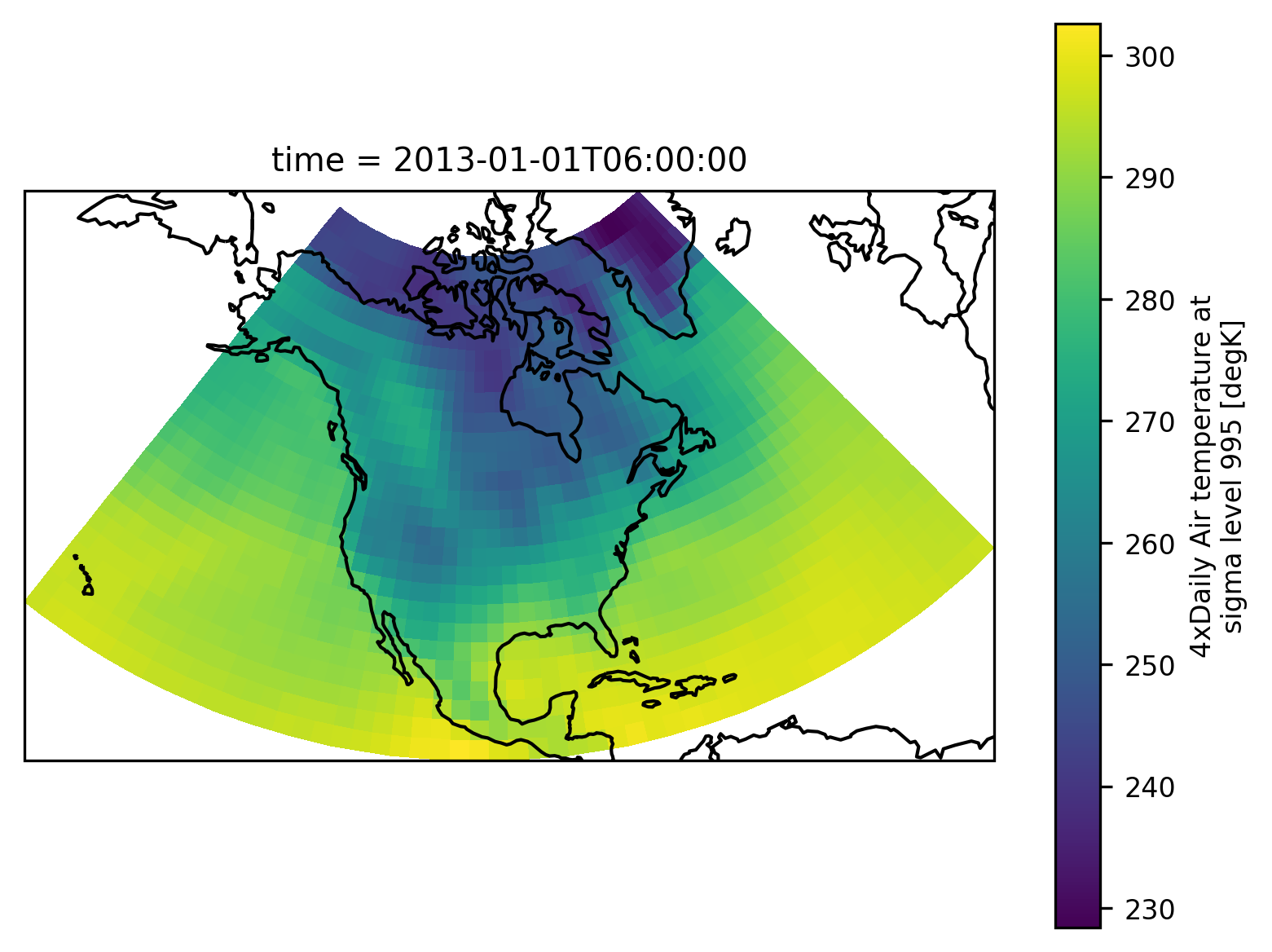
An xarray object can be passed to the mask_3D function:
mask_3D = regionmask.defined_regions.srex.mask_3D(airtemps)
mask_3D
<xarray.DataArray 'mask' (region: 6, lat: 25, lon: 53)> Size: 8kB
False False False False False ... False False False False
Coordinates:
* lat (lat) float32 100B 75.0 72.5 70.0 ... 17.5 15.0
* lon (lon) float32 212B 200.0 202.5 ... 327.5 330.0
* region (region) int64 48B 1 2 3 4 5 6
abbrevs (region) <U3 72B 'ALA' 'CGI' ... 'ENA' 'CAM'
names (region) <U22 528B 'Alaska/N.W. Canada' ... ...
Attributes:
standard_name: regionPer default this creates a mask containing one layer (slice) for
each region containing (at least) one gridpoint. As the example data
only has values over Northern America we only get only 6 layers even
though there are 26 SREX regions. To obtain all layers specify
drop=False:
mask_full = regionmask.defined_regions.srex.mask_3D(airtemps, drop=False)
mask_full
<xarray.DataArray 'mask' (region: 26, lat: 25, lon: 53)> Size: 34kB
False False False False False ... False False False False
Coordinates:
* lat (lat) float32 100B 75.0 72.5 70.0 ... 17.5 15.0
* lon (lon) float32 212B 200.0 202.5 ... 327.5 330.0
* region (region) int64 208B 1 2 3 4 5 ... 23 24 25 26
abbrevs (region) <U3 312B 'ALA' 'CGI' ... 'NAU' 'SAU'
names (region) <U24 2kB 'Alaska/N.W. Canada' ... '...
Attributes:
standard_name: regionNote mask_full now has 26 layers.
Select a region
As mask_3D contains region, abbrevs, and names as
(non-dimension) coordinates we can use each of those to select an
individual region:
# 1) by the index of the region:
r1 = mask_3D.sel(region=3)
# 2) with the abbreviation
r2 = mask_3D.isel(region=(mask_3D.abbrevs == "WNA"))
# 3) with the long name:
r3 = mask_3D.isel(region=(mask_3D.names == "E. North America"))
This also applies to the regionally-averaged data below.
It is currently not possible to use sel with a non-dimension
coordinate - to directly select abbrev or name you need to
create a MultiIndex:
mask_3D.set_index(regions=["region", "abbrevs", "names"]);
Mask out a region
Using where a specific region can be ‘masked out’ (i.e. all data
points outside of the region become NaN):
airtemps_cna = airtemps.where(r1)
Which looks as follows:
proj = ccrs.LambertConformal(central_longitude=-100)
ax = plt.subplot(111, projection=proj)
airtemps_cna.isel(time=1).air.plot(ax=ax, transform=ccrs.PlateCarree())
ax.coastlines();
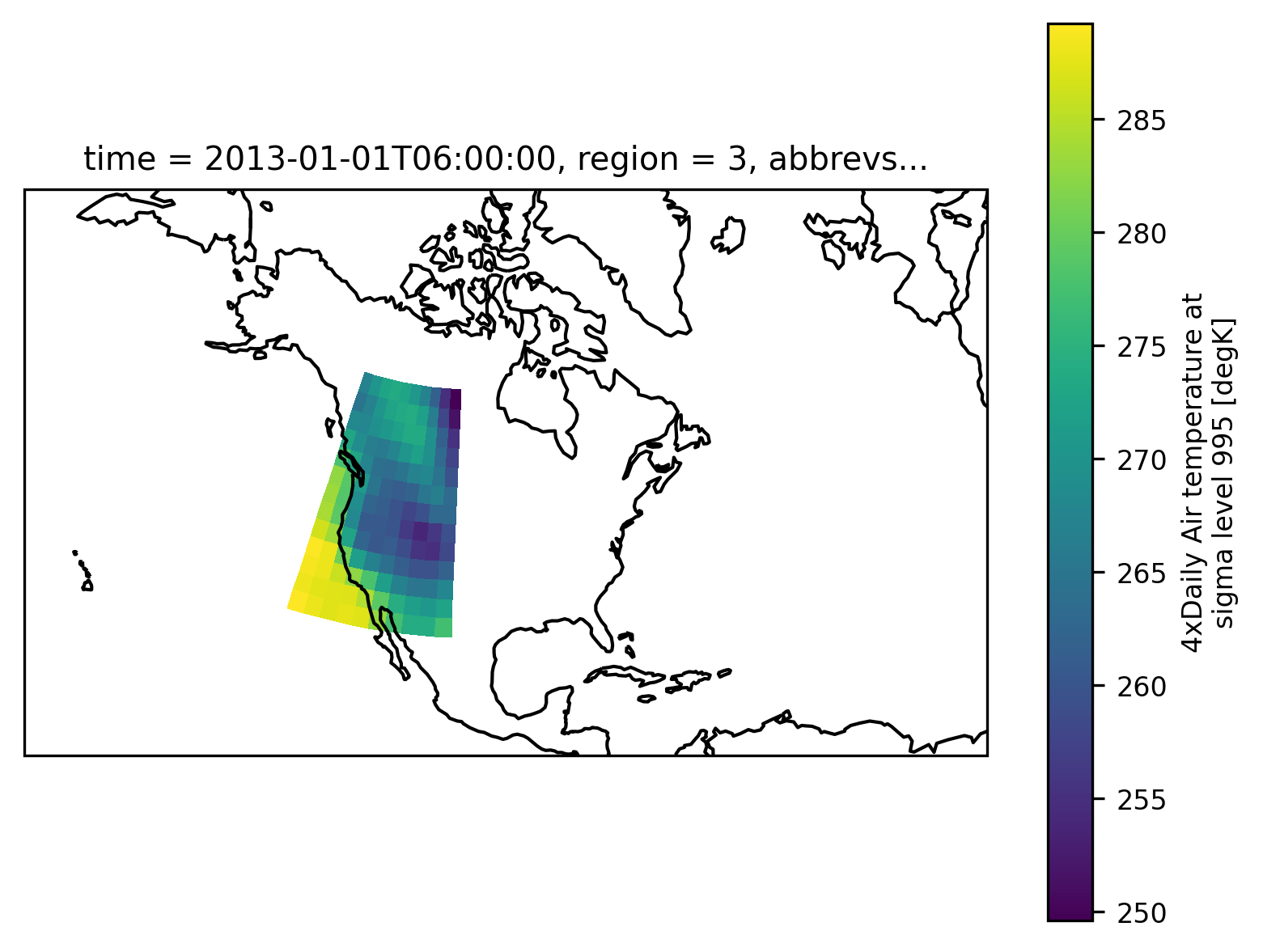
We could now use airtemps_cna to calculate the regional average for
‘Central North America’. However, there is a more elegant way.
Calculate weighted regional averages
Using the 3-dimensional mask it is possible to calculate weighted
averages of all regions in one go, using the weighted method
(requires xarray 0.15.1 or later). As proxy of the grid cell area we use
cos(lat).
Note
It is better to use a model’s original grid cell area (e.g. areacella). cos(lat) works reasonably well for regular lat/ lon grids. For irregular grids (regional models, ocean models, …) it is not appropriate.
weights = np.cos(np.deg2rad(airtemps.lat))
ts_airtemps_regional = airtemps.weighted(mask_3D * weights).mean(dim=("lat", "lon"))
Let’s break down what happens here. By multiplying mask_3D * weights
we get a DataArray where gridpoints not in the region get a weight of 0.
Gridpoints within a region get a weight proportional to the gridcell
area. airtemps.weighted(mask_3D * weights) creates an xarray object
which can be used for weighted operations. From this we calculate the
weighted mean over the lat and lon dimensions. The resulting
dataarray has the dimensions region x time:
ts_airtemps_regional
<xarray.Dataset> Size: 94kB
Dimensions: (time: 2920, region: 6)
Coordinates:
* time (time) datetime64[ns] 23kB 2013-01-01 ... 20...
* region (region) int64 48B 1 2 3 4 5 6
abbrevs (region) <U3 72B 'ALA' 'CGI' ... 'ENA' 'CAM'
names (region) <U22 528B 'Alaska/N.W. Canada' ... ...
Data variables:
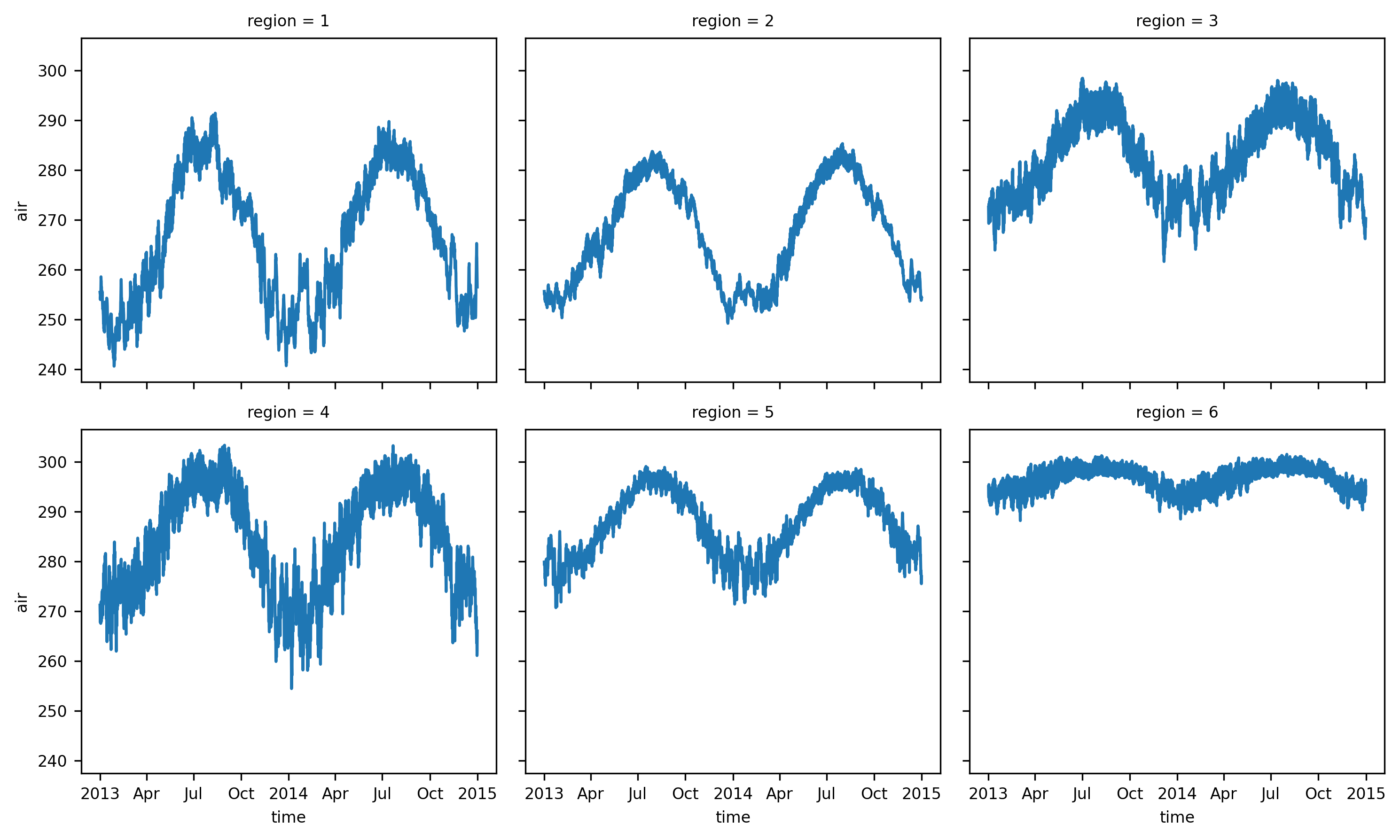
air (time, region) float32 70kB 255.5 ... 295.0The regionally-averaged time series can be plotted:
ts_airtemps_regional.air.plot(col="region", col_wrap=3);
Restrict the mask to land points
Combining the mask of the regions with a land-sea mask we can create a
land-only mask using the land_110 region from NaturalEarth.
Warning
It is better to use a model’s original land/ sea mask (e.g. sftlf). Many CMIP models treat the Antarctic ice shelves and the Caspian Sea as land, while it is classified as ‘water’ in natural_earth_v5_0_0.land_110.
With this caveat in mind we can create the land-sea mask:
land_110 = regionmask.defined_regions.natural_earth_v5_0_0.land_110
land_mask = land_110.mask_3D(airtemps)
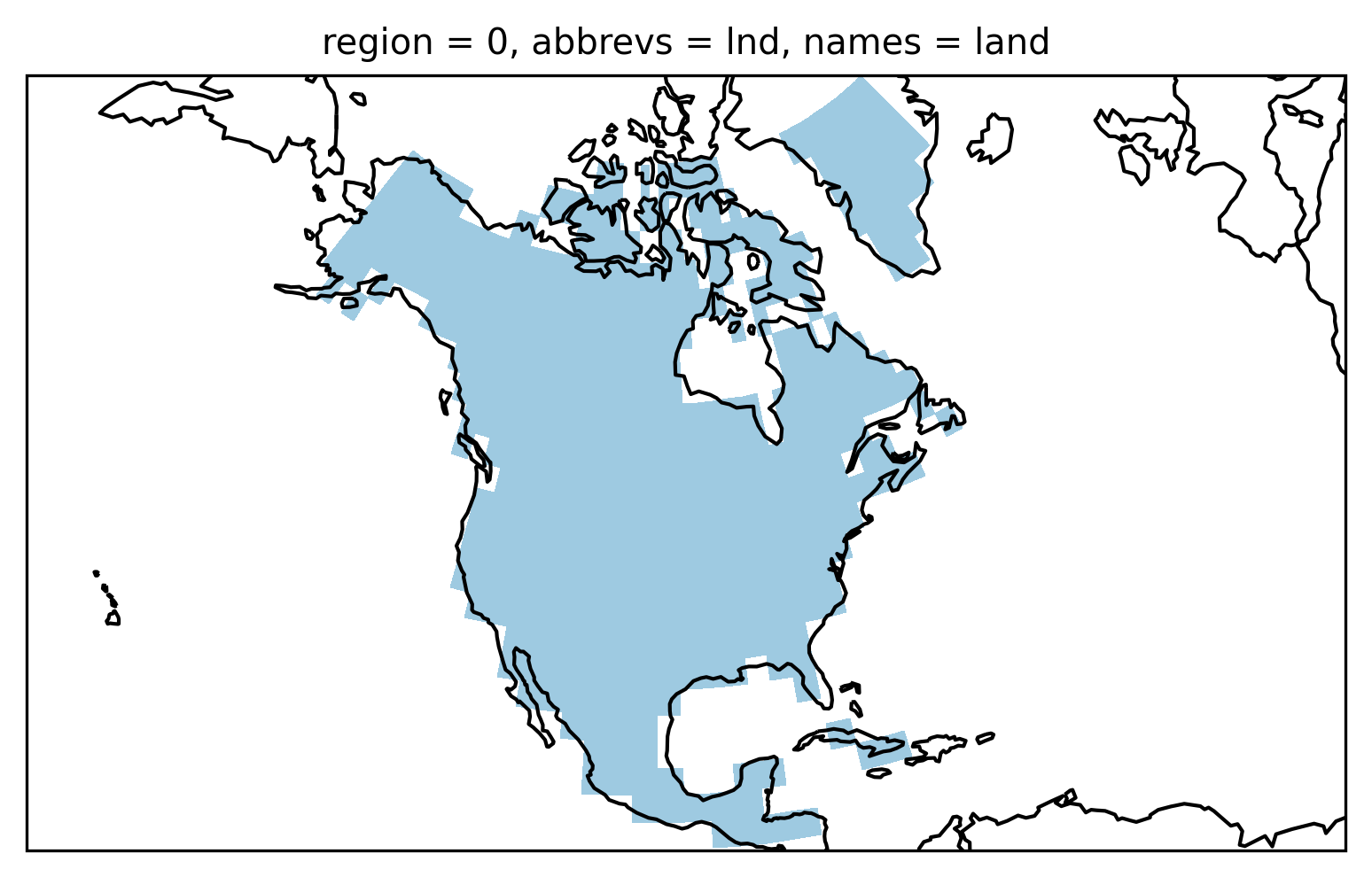
and plot it
proj = ccrs.LambertConformal(central_longitude=-100)
ax = plt.subplot(111, projection=proj)
land_mask.squeeze().plot.pcolormesh(
ax=ax, transform=ccrs.PlateCarree(), cmap=cmap1, add_colorbar=False
)
ax.coastlines();
To create the combined mask we multiply the two:
mask_lsm = mask_3D * land_mask.squeeze(drop=True)
Note the .squeeze(drop=True). This is required to remove the
region dimension from land_mask.
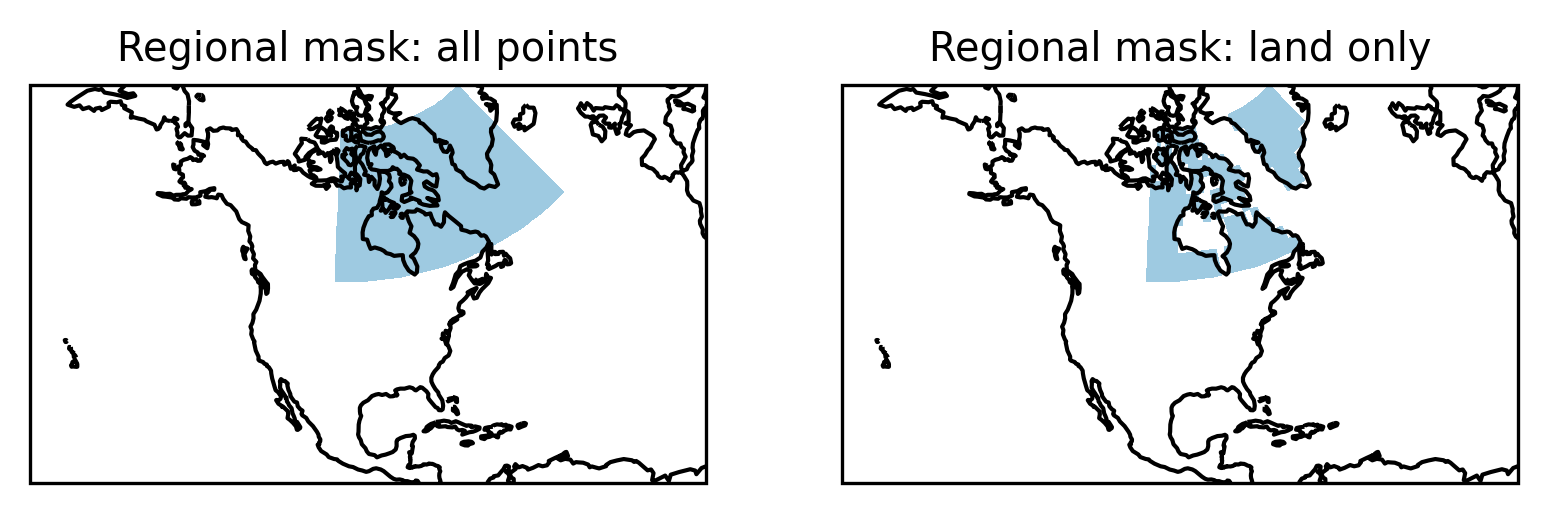
Finally, we compare the original mask with the one restricted to land points:
f, axes = plt.subplots(1, 2, subplot_kw=dict(projection=proj))
ax = axes[0]
mask_3D.sel(region=2).plot(
ax=ax, transform=ccrs.PlateCarree(), add_colorbar=False, cmap=cmap1
)
ax.coastlines()
ax.set_title("Regional mask: all points")
ax = axes[1]
mask_lsm.sel(region=2).plot(
ax=ax, transform=ccrs.PlateCarree(), add_colorbar=False, cmap=cmap1
)
ax.coastlines()
ax.set_title("Regional mask: land only");
References
Special Report on Managing the Risks of Extreme Events and Disasters to Advance Climate Change Adaptation (SREX, Seneviratne et al., 2012)